參考網站
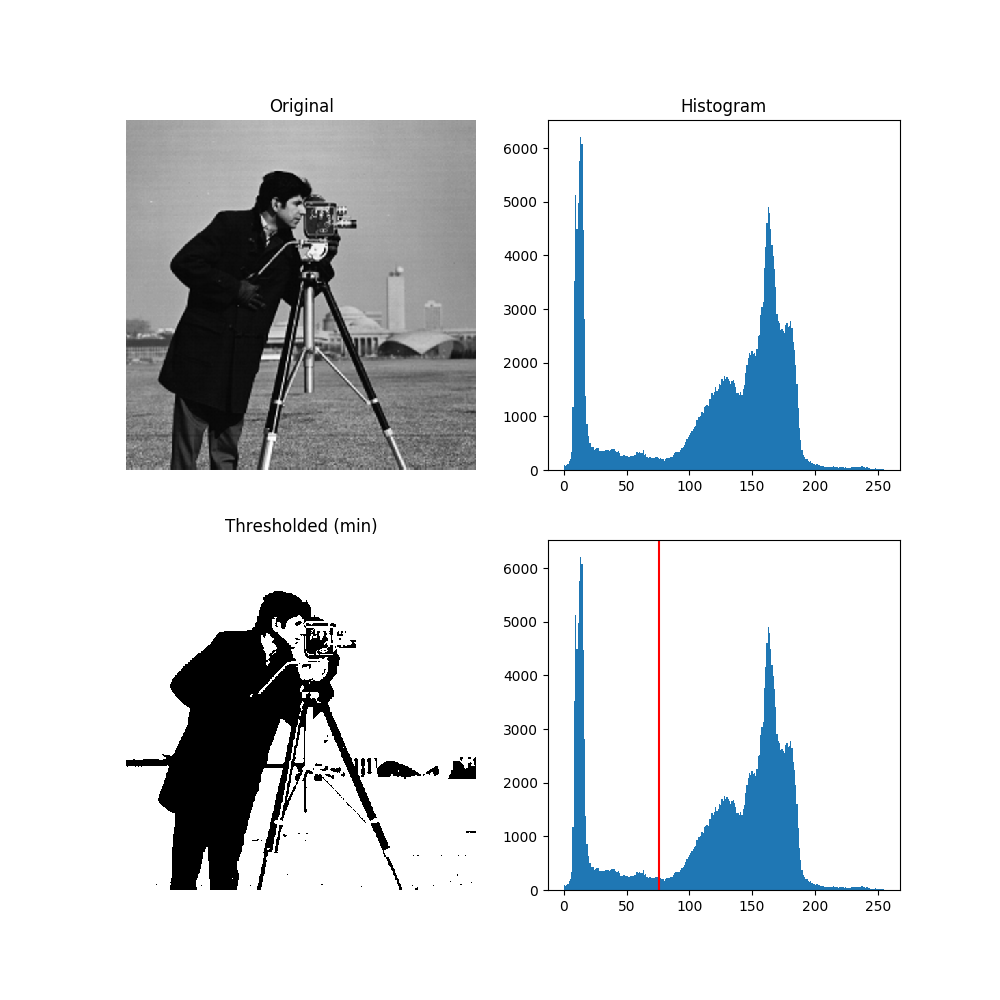
當只想要物體的形狀或面積,可先將histogram分成兩個值0(白)與255(黑)方便後續的輪廓描繪與計算。
這邊來學習scikit-image上面的例子:
這裡測試
import matplotlib.pyplot as plt
from skimage import data
from skimage.filters import try_all_threshold
img = data.page()
fig, ax = try_all_threshold(img, figsize=(10, 8), verbose=False)
plt.show()

from skimage.filters import threshold_mean
image = data.camera()
thresh = threshold_mean(image)
binary = image > thresh
fig, axes = plt.subplots(ncols=2, figsize=(8, 3))
ax = axes.ravel()
ax[0].imshow(image, cmap=plt.cm.gray)
ax[0].set_title('Original image')
ax[1].imshow(binary, cmap=plt.cm.gray)
ax[1].set_title('Result')
for a in ax:
a.axis('off')
plt.show()
增加local threshold的設定,補足左半部較暗區的文字與白紙的區隔。
from skimage.filters import threshold_otsu, threshold_local
image = data.page()
global_thresh = threshold_otsu(image)
binary_global = image > global_thresh
block_size = 35
local_thresh = threshold_local(image, block_size, offset=10)
binary_local = image > local_thresh
fig, axes = plt.subplots(nrows=3, figsize=(7, 8))
ax = axes.ravel()
plt.gray()
ax[0].imshow(image)
ax[0].set_title('Original')
ax[1].imshow(binary_global)
ax[1].set_title('Global thresholding')
ax[2].imshow(binary_local)
ax[2].set_title('Local thresholding')
for a in ax:
a.axis('off')
plt.show()

如何擷取物體邊緣的輪廓在鄰域處理也是很常用的方法,從youtube的動圖可已清楚了解mask矩陣是怎麼作用的。接下來測試scikit-image裡面幾個演算法範例:
import numpy as np
import matplotlib.pyplot as plt
from skimage.data import camera
from skimage.filters import roberts, sobel, scharr, prewitt
image = camera()
edge_roberts = roberts(image)
edge_sobel = sobel(image)
fig, ax = plt.subplots(ncols=2, sharex=True, sharey=True,
figsize=(8, 4))
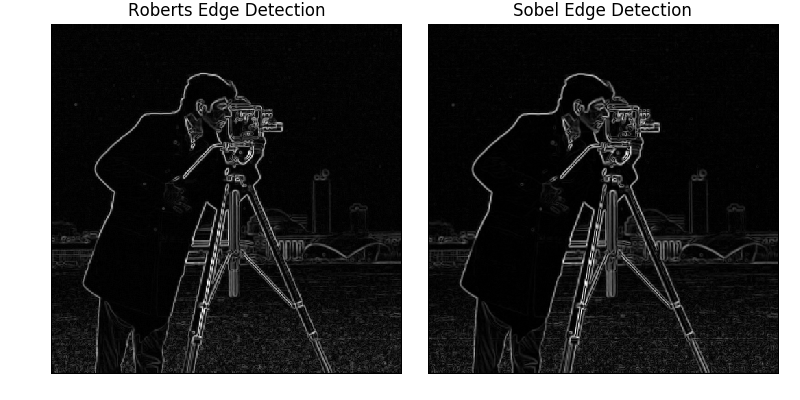
ax[0].imshow(edge_roberts, cmap=plt.cm.gray)
ax[0].set_title('Roberts Edge Detection')
ax[1].imshow(edge_sobel, cmap=plt.cm.gray)
ax[1].set_title('Sobel Edge Detection')
for a in ax:
a.axis('off')
plt.tight_layout()
plt.show()

import numpy as np
import matplotlib.pyplot as plt
from scipy import ndimage as ndi
from skimage import feature
# Generate noisy image of a square
im = np.zeros((128, 128))
im[32:-32, 32:-32] = 1
im = ndi.rotate(im, 15, mode='constant')
im = ndi.gaussian_filter(im, 4)
im += 0.2 * np.random.random(im.shape)
# Compute the Canny filter for two values of sigma
edges1 = feature.canny(im)
edges2 = feature.canny(im, sigma=3)
# display results
fig, (ax1, ax2, ax3) = plt.subplots(nrows=1, ncols=3, figsize=(8, 3),
sharex=True, sharey=True)
ax1.imshow(im, cmap=plt.cm.gray)
ax1.axis('off')
ax1.set_title('noisy image', fontsize=20)
ax2.imshow(edges1, cmap=plt.cm.gray)
ax2.axis('off')
ax2.set_title('Canny filter, $\sigma=1$', fontsize=20)
ax3.imshow(edges2, cmap=plt.cm.gray)
ax3.axis('off')
ax3.set_title('Canny filter, $\sigma=3$', fontsize=20)
fig.tight_layout()
plt.show()
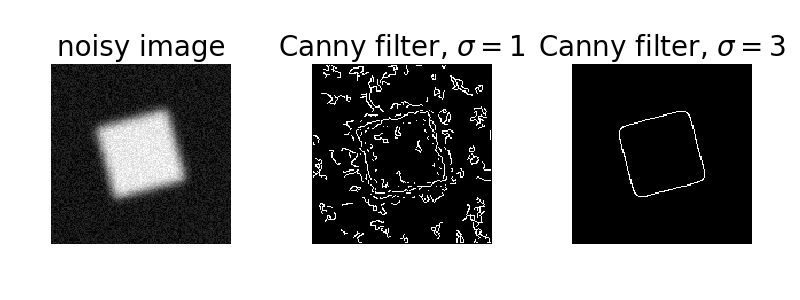

Canny 這個演算法用了多次的邊緣擷取函數,還可將雜質濾除~
